Research
|
Anthropogenic climate change is now recognized as a major force driving alterations in the fitness, behaviour, distribution and ecology of species across the planet. Climate exerts powerful effects on the distribution and abundance of the earth's insect species. Based on major climatic models Earth’s temperature is going to rise 3-6 °C by the end of this century and this climate change related warming is going to generate changes for many insect populations and the ecosystems they inhabit. The unprecedented rates of climate changes in the future, coupled with land use changes that impede gene flow, can be expected to disrupt the entire ecology of many species. Warmer temperatures, and desertification associated with climate changes will tend to influence (and frequently amplify) insect species’ population dynamics directly through effects on survival, generation time, fecundity, response to stress and dispersal. A systematic and in-depth approach provides a foundation for describing how insect species are responding to recent climatic trends on the basis of insect physiology, and predicting generalized species distributions and population dynamics for the future.
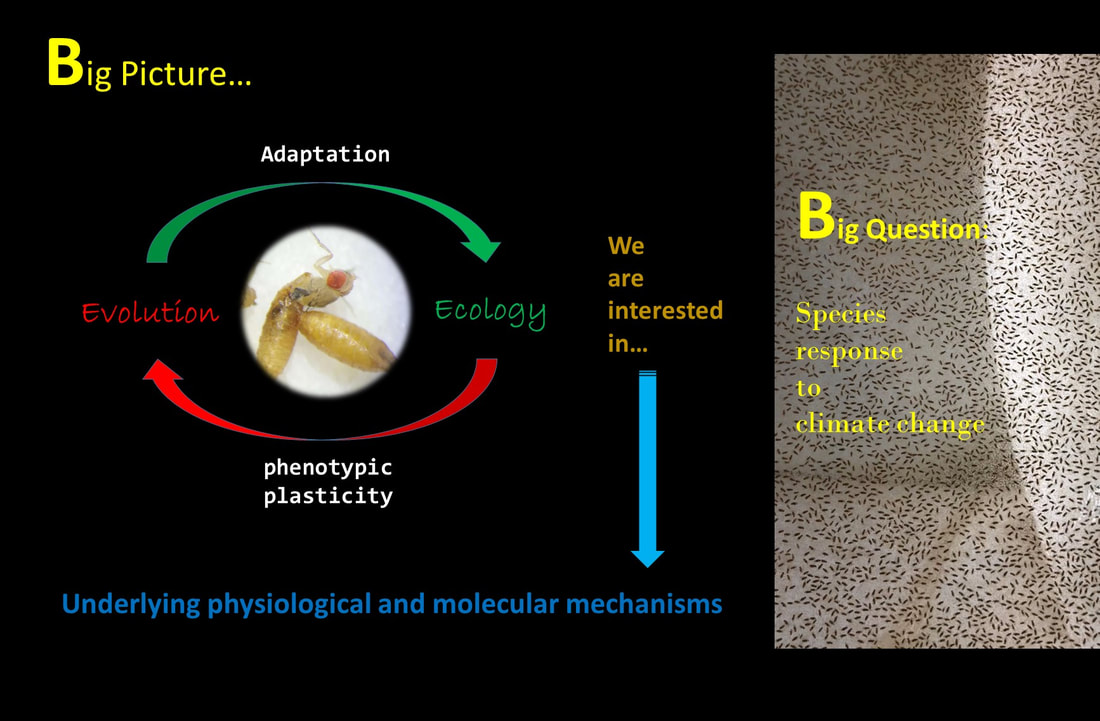
EEE Lab is interested in understanding responses of organisms to changing environment i.e. warming. Our approach is from ‘macrophysiology to molecules’. The focus is on interaction between phenotypic plasticity and adaptation, and relationship between underlying physiological and molecular mechanisms. Ecological changes drive evolution which acts on organism’s physiology and thereby fitness. The ecological conditions are never stable, and organisms must cope with varying conditions i.e. physiological performance or some form of regulation. We do it by either manipulating organisms or their environments. We also make use of experimental evolution to study how physiological systems function and evolve under defined conditions. The combined approach we follow helps us understand ecological patterns and processes, survival in and adaptation to a changing world.
EEE Lab also hosts a resource on Indian Drosophila Ecology & Evolution (a window to Indian Drosophila clines DrosoCline). The research findings associated to this long-term study are regularly updated on the following web-resource:
http://www.rajpurohit-lab.org/drosocline.html
Ongoing Projects:
1. Genetic background (alleles) and evolutionary potential of natural populations
2. Cuticular hydrocarbon and melanin interactions in natural populations
3. Artificial selection (ecologically relevant traits)
3. Impact of drought (water) and temperature (heat) on mating behavior
In Focus:
I. Spatiotemporal variations as a tool to understand organismal responses to climate change:
In the past, we studied ecologically relevant traits in Drosophila species populations along spatial and temporal scales in India (reviewed in Rajpurohit et al. 2017). This work established that seasonal temperature variability along the Indian latitudes play a critical role in defining various fitness trait clines (i.e. an increase or decrease in a trait value along the latitudes or altitude). We are trying to find out the genetic architecture of populations collected along the latitudes and altitudes. We use high-throughput platforms/approaches-genome resequencing of natural populations, gene expression analysis, and metabolomics to understand the molecular genetic wiring of natural populations. The efforts are in direction to integrate findings from genomics and metabolomics to understand the organismal responses to climate change.
In the past, we studied ecologically relevant traits in Drosophila species populations along spatial and temporal scales in India (reviewed in Rajpurohit et al. 2017). This work established that seasonal temperature variability along the Indian latitudes play a critical role in defining various fitness trait clines (i.e. an increase or decrease in a trait value along the latitudes or altitude). We are trying to find out the genetic architecture of populations collected along the latitudes and altitudes. We use high-throughput platforms/approaches-genome resequencing of natural populations, gene expression analysis, and metabolomics to understand the molecular genetic wiring of natural populations. The efforts are in direction to integrate findings from genomics and metabolomics to understand the organismal responses to climate change.
II. Evolutionary potential of populations/species (Field and Laboratory conditions)
There is a possibility that if populations are not adapting fast enough in the changing climatic conditions they might go extinct. At this important juncture we might need to dig biological complexities a bit deeper and explore the capacities of natural populations to adapt genetically to environmental changes. We expose known populations to growing tropical summers and check their genetic capacities. This approach helps us to test various ecological hypotheses. Experimental Evolution Study Stations at the Ahmedabad University has a set of mesocosm units to test such hypotheses (see Rajpurohit et al. 2018). This set-up is equipped with a climate tower to collect weather data day/night to keep track of population’s responses to climatic variables.
There is a possibility that if populations are not adapting fast enough in the changing climatic conditions they might go extinct. At this important juncture we might need to dig biological complexities a bit deeper and explore the capacities of natural populations to adapt genetically to environmental changes. We expose known populations to growing tropical summers and check their genetic capacities. This approach helps us to test various ecological hypotheses. Experimental Evolution Study Stations at the Ahmedabad University has a set of mesocosm units to test such hypotheses (see Rajpurohit et al. 2018). This set-up is equipped with a climate tower to collect weather data day/night to keep track of population’s responses to climatic variables.
Current Collaborators:
- Hosken Lab (Exeter University, Penryn, UK)
- Loeschcke Lab (Aarhus University, Aarhus, Denmark)
- Merritt Lab (Laurentian University, Sudbury, Canada)
- Gibbs Lab (University of Nevada, Las Vegas, USA)
- Pawar Lab (IMPERIAL UK)
- Schmidt Lab (University of Pennsylvania, Philadelphia, USA)